Making sense of atomic design: molecules and organisms
Our Interaction Designer, Alla Kholmatova, reflects on some of the aspects of using atomic design at FutureLearn.

A year ago we wrote about building our first pattern library at FutureLearn and why we decided to use atomic design methodology. Overall, atomic design has worked out well for our team. It has helped us to build a shared understanding of the interface, make a transition towards a more modular system, and evolve our design language.
While some concepts in atomic design were clear from the start, others felt a bit foggy. For example, one area that we’re still trying to improve our understanding of, is the difference between molecules and organisms.
Molecules and organisms
Brad Frost defines molecules as “groups of atoms bonded together, the smallest fundamental units of a compound”, and organisms as “groups of molecules joined together to form a relatively complex, distinct section of an interface”.
While these definitions may sound simple in theory, we’ve been struggling to understand exactly when a molecule is complex enough to become an organism, the defining difference between the two types, and even why we need both types in the first place. (The CSS part of organisms and molecules are quite similar, wouldn’t it be simpler to just have smaller and larger molecules?)
Why have both types in the first place?
We felt that a separation between molecules and organisms brings (or could potentially bring) three main advantages.
- From the front-end perspective, organisms are good candidates for separating their HTML parts into partials (molecules are probably too simple for this).
- From the design perspective, it helps to understand the design, and see what parts it consists of. It also helps to find the right element quicker to use in your design (for example, “I need a supporting element here to enable sharing of this content, let’s see what we have…”)
- From the Pattern Library perspective, it helps to organise the elements and makes them easier to find (at least easier than the one long list we used to have).
The problem we were trying to address
However, there was no shared understanding in the team about what the distinction is exactly. Is it about the visual presentation – for example, size or visual complexity? Is it about the complexity or importance of the content? Is it about the function? What if the content alters the function? And what if the function is relative, depending on the context in which an element appears?
As a result, we used to spend a long time debating whether something is a molecule or an organism. The debates weren’t always productive, and we couldn’t afford to continue having them.
So we decided to run a workshop with designers and front-end developers, to see how we could become more efficient in classifying the elements.
The workshop
As always, we cut the interface elements out on paper (pictured above). We split into small teams and asked each group to sort the interface elements into groups (similar to a card sort), in the way that made sense to them, rather than the way they were already organised in the pattern library. The idea was just to see what groupings people would come up with.
The results
The results of the workshop were fairly inconclusive. The teams classified the modules in a variety of ways and ultimately ended up with different groupings.
However, there were clearly two distinct types. We all picked up that some modules could be used on their own, while others could only work only as part of another module. For example, these two components are quite different, both on the functional and presentation levels:
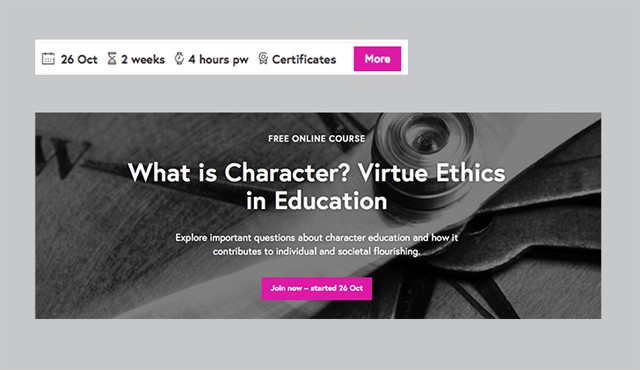
The elements in the first group (which the top component represents) have more of a supporting function – let’s call them “helpers”. The elements in the second group are more independent – let’s call them “standalone”.
Helpers
Helpers don’t make sense on their own. Here are just a few typical representatives of this group. In our interface molecules are typically “helpers”.
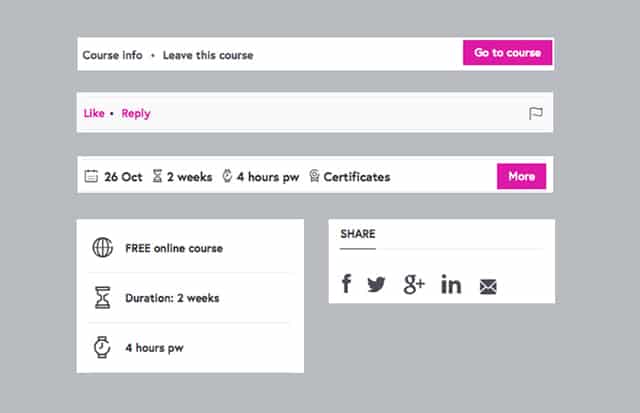
Standalones
The “standalone” elements can “survive” on their own – they can be viewed as their own self-contained units of content and functionality. Typical representatives of standalone modules on FutureLearn are:
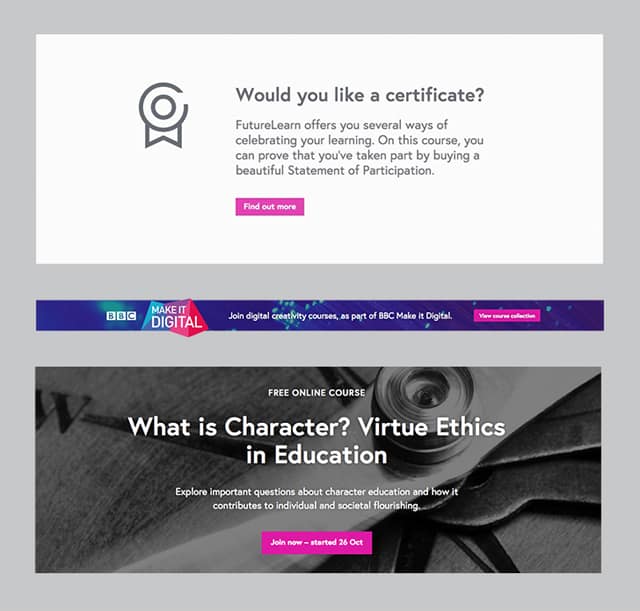
In our interface, organisms are typically “standalone”.
So far it makes sense. Molecules primarily support other interface elements. Organisms are primarily self contained and can stand on their own.
Helper or standalone?
However, many elements fall somewhere in between. The ones that fall in between can in theory belong to either group, depending on two main factors – content and context. Content and context are of course subject to change, which makes it difficult to classify those elements. Here is a typical example of such module:
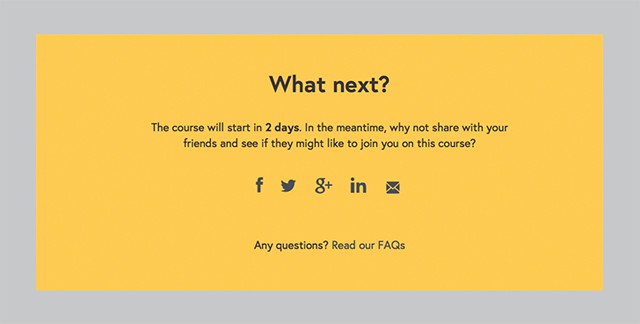
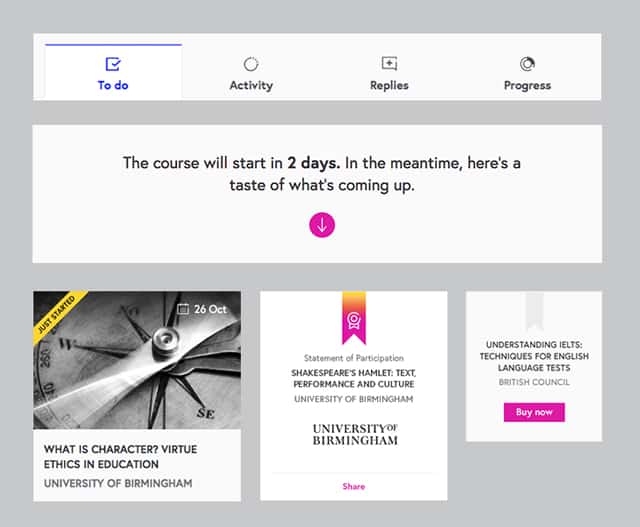
On the surface this looks like an organism (it’s a fairly complex element which contains a fair bit of content). However, it can’t really stand on its own. It’s not clear what exactly you’re sharing, and which course will start in 2 days. But imagine that we changed the content, gave it a more meaningful title, and referenced the title of the course. Suddenly this module can stand on its own.
Our interface abounds with elements like those, that don’t fall clearly in one of those two buckets. Here’re just a few of them:
The takeaways
Even after a year of working with atomic design there is still no clarity and shared understanding in our team about how interface elements should be classified. But maybe it’s OK. There isn’t always the “right” way to classify and organise things.
One thing we learned is that we should be more flexible in deciding which group a module belongs to. Rather than involving the whole team in the debate, the designer and developer who introduce a new module ultimately should decide whether it is a molecule or an organism. The rest of the team should accept their decision, unless they have strong, well-grounded objections.
When thinking about complexity of elements, it helps viewing molecules as “helpers” and organisms as “standalone” modules. If an element doesn’t clearly fall into either group, asking these questions may help:
- Is this more of a supporting element or a standalone one?
- Would it normally be part of something else and be reused within various components? (probably molecule)
- Is it a well defined and relatively independent part or section of a page? (probably organism)
If you work with atomic design, we’d love to hear your thoughts. Do you see the value in separating molecules from organisms? If so, how do you distinguish the two? Tell us in the comments below. You might also be interested in design roles at FutureLearn.




